Cancer genomes contain large numbers of somatic alterations but few genes drive tumor development. Identifying cancer driver genes is critical for precision oncology. Most of current approaches either identify driver genes based on mutational recurrence or using estimated scores predicting the functional consequences of mutations.
driveR is a tool for personalized or batch analysis of genomics data for driver gene prioritization by combining genomics information and prior biological knowledge. As features, driveR uses coding impact metaprediction scores, non-coding impact scores, somatic copy number alteration scores, hotspot gene/double-hit gene condition, ‘phenolyzer’ gene scores and memberships to cancer-related KEGG pathways. It uses these features to estimate cancer-type-specific probabilities for each gene of being a cancer driver using the related task of a multi-task learning classification model.
The method is described in detail in Ülgen E, Sezerman OU. driveR: a novel method for prioritizing cancer driver genes using somatic genomics data. BMC Bioinformatics. 2021 May 24;22(1):263.https://doi.org/10.1186/s12859-021-04203-7
Installation
You can install the latest released version of driveR from CRAN via:
install.packages("driveR")You can install the development version of driveR from GitHub with:
# install.packages("devtools")
devtools::install_github("egeulgen/driveR", build_vignettes = TRUE)Usage
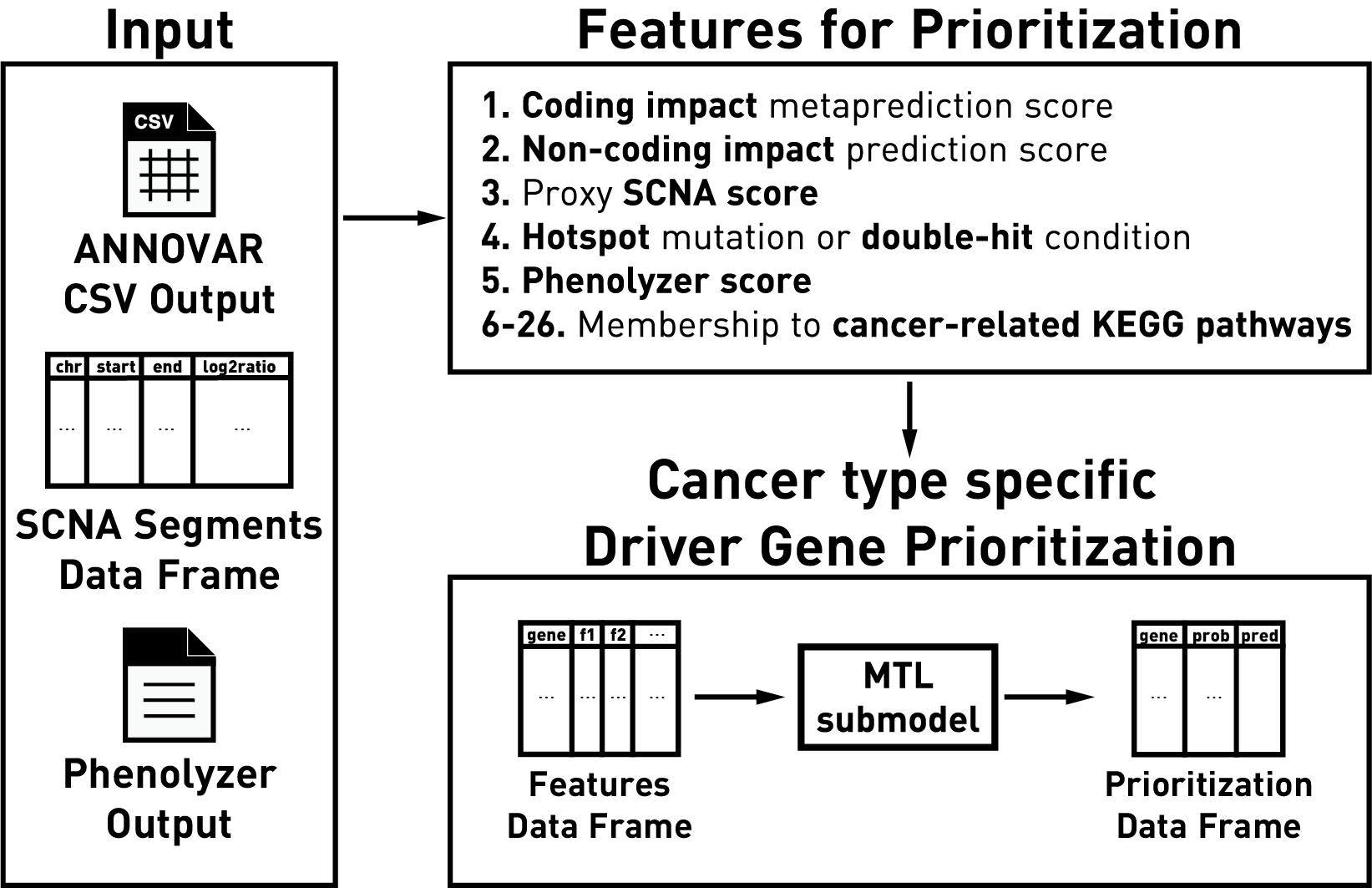
driveR has two main objectives:
- Prediction of impact of coding variants (achieved via
predict_coding_impact()) -
Prioritization of cancer driver genes (achieved via
create_features_df()andprioritize_driver_genes())
Note that driveR require operations outside of R and depends on the outputs from the external tools ANNOVAR and phenolyzer.
For detailed information on how to use driveR, please see the vignette “How to use driveR” via vignette("how_to_use")
 driveR: An R Package for Prioritizing Cancer Driver Genes Using Genomics Data
driveR: An R Package for Prioritizing Cancer Driver Genes Using Genomics Data