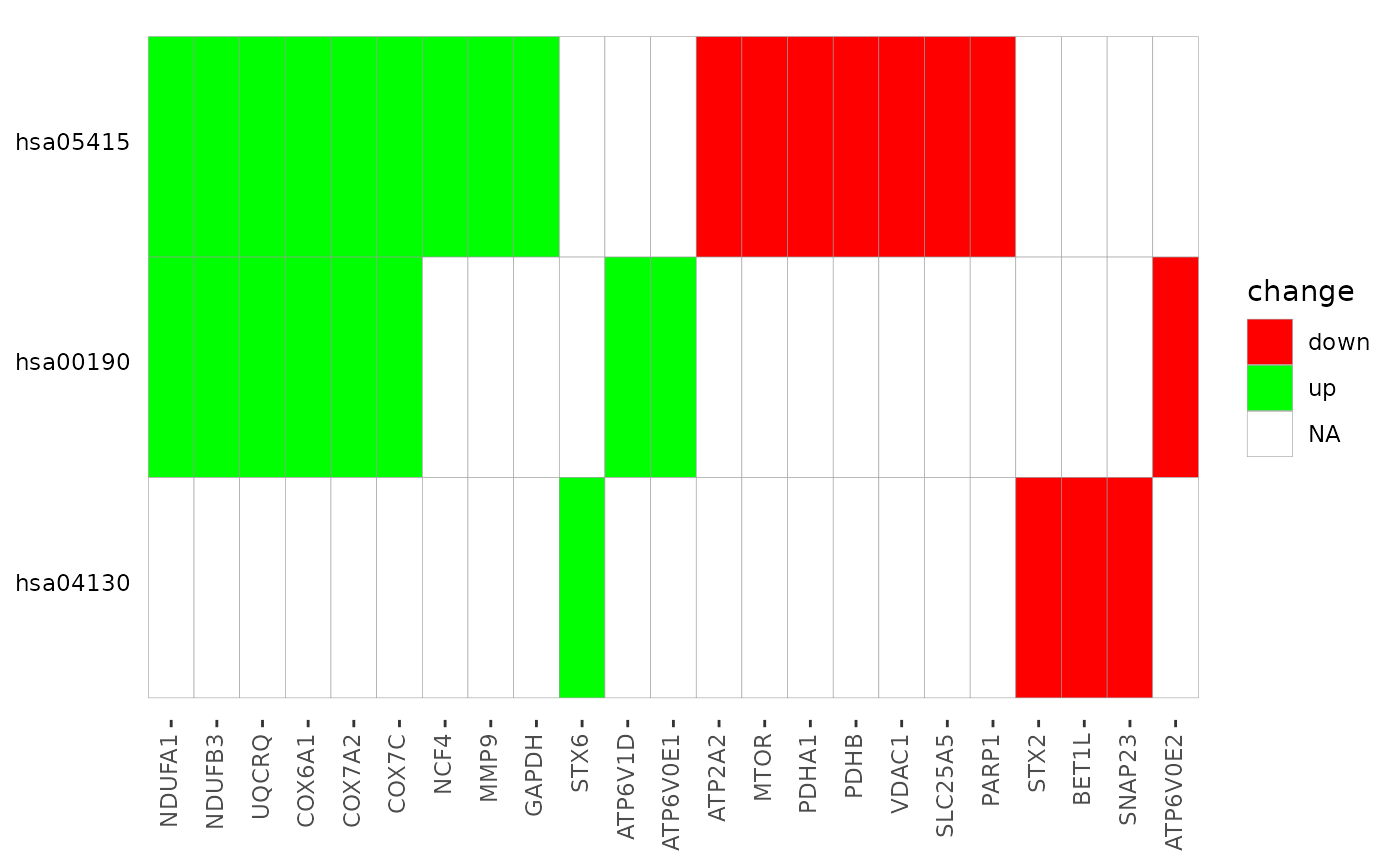
Create Terms by Genes Heatmap
term_gene_heatmap(
result_df,
genes_df,
num_terms = 10,
use_description = FALSE,
low = "red",
mid = "black",
high = "green",
legend_title = "change",
sort_terms_by_p = FALSE,
...
)Arguments
- result_df
A dataframe of pathfindR results that must contain the following columns:
- Term_Description
Description of the enriched term (necessary if
use_description = TRUE)- ID
ID of the enriched term (necessary if
use_description = FALSE)- lowest_p
the highest adjusted-p value of the given term over all iterations
- Up_regulated
the up-regulated genes in the input involved in the given term's gene set, comma-separated
- Down_regulated
the down-regulated genes in the input involved in the given term's gene set, comma-separated
- genes_df
the input data that was used with
run_pathfindR. It must be a data frame with 3 columns:Gene Symbol (Gene Symbol)
Change value, e.g. log(fold change) (optional)
p value, e.g. adjusted p value associated with differential expression
The change values in this data frame are used to color the affected genes
- num_terms
Number of top enriched terms to use while creating the plot. Set to
NULLto use all enriched terms (default = 10)- use_description
Boolean argument to indicate whether term descriptions (in the 'Term_Description' column) should be used. (default =
FALSE)- low
a string indicating the color of 'low' values in the coloring gradient (default = 'green')
- mid
a string indicating the color of 'mid' values in the coloring gradient (default = 'black')
- high
a string indicating the color of 'high' values in the coloring gradient (default = 'red')
- legend_title
legend title (default = 'change')
- sort_terms_by_p
boolean to indicate whether to sort terms by 'lowest_p' (
TRUE) or by number of genes (FALSE) (default =FALSE)- ...
additional arguments for
input_processing(used ifgenes_dfis provided)
Value
a ggplot2 object of a heatmap where rows are enriched terms and
columns are involved input genes. If genes_df is provided, colors of
the tiles indicate the change values.
Examples
term_gene_heatmap(example_pathfindR_output, num_terms = 3)