Overview
pathfindR is an R package for enrichment analysis via active subnetworks. The package also offers functionality to cluster the enriched terms and identify representative terms in each cluster, score the enriched terms per sample, and visualize analysis results. As of the latest version, the package also allows comparison of two pathfindR results.
The functionality suite of pathfindR is described in detail in Ulgen E, Ozisik O, Sezerman OU. 2019. pathfindR: An R Package for Comprehensive Identification of Enriched Pathways in Omics Data Through Active Subnetworks. Front. Genet. https://doi.org/10.3389/fgene.2019.00858
For detailed documentation, see pathfindR’s website.
Installation
- You can install the released version of pathfindR from CRAN via:
install.packages("pathfindR")- Since version 2.1.0, you may also install
pathfindRvia conda:
- Via pak (this might be preferable given
pathfindR’s Bioconductor dependencies):
install.packages("pak") # if you have not installed "pak"
pak::pkg_install("pathfindR")- And the development version from GitHub via
devtools:
install.packages("devtools") # if you have not installed "devtools"
devtools::install_github("egeulgen/pathfindR")IMPORTANT NOTE For the active subnetwork search component to work, the user must have Java (>= 8.0) installed, and the path/to/java must be in the PATH environment variable.
We also have docker images available on Docker Hub and GitHub:
# pull image for the latest release
docker pull egeulgen/pathfindr:latest
# pull image for a specific version (e.g., 1.4.1)
docker pull egeulgen/pathfindr:1.4.1Online app on superbio.ai: https://app.superbio.ai/apps/111/
Enrichment Analysis with pathfindR
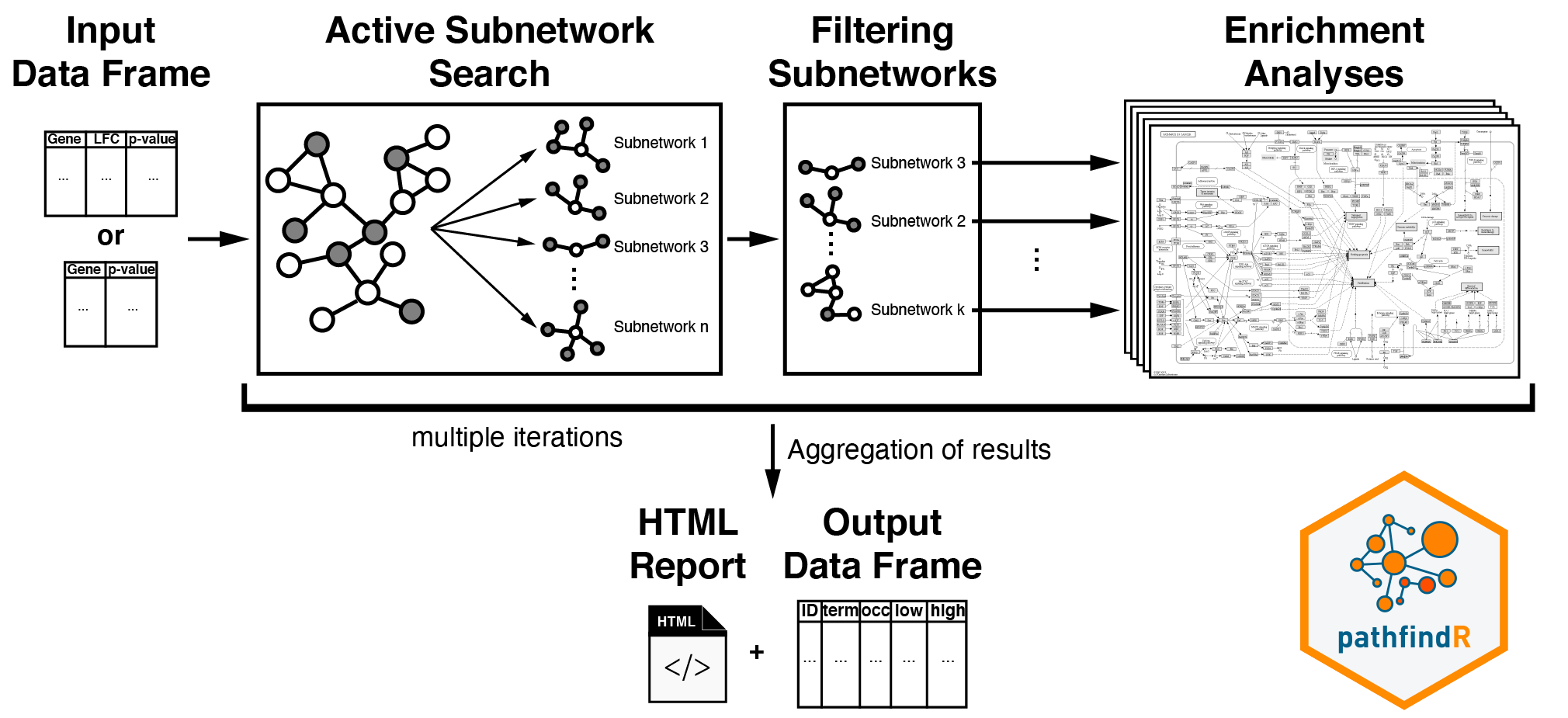
This workflow takes in a data frame consisting of “gene symbols”, “change values” (optional), and “associated p-values”:
| Gene_symbol | logFC | FDR_p |
|---|---|---|
| FAM110A | -0.69 | 3.4e-06 |
| RNASE2 | 1.35 | 1.0e-05 |
| S100A8 | 1.54 | 3.5e-05 |
| S100A9 | 1.03 | 2.3e-04 |
After input testing, any gene symbol that is not in the chosen protein-protein interaction network (PIN) is converted to an alias symbol if there is an alias that is found in the PIN. After mapping the input genes with the associated p-values onto the PIN, active subnetwork search is performed. The resulting active subnetworks are then filtered based on their scores and the number of significant genes they contain.
An active subnetwork can be defined as a group of interconnected genes in a protein-protein interaction network (PIN) that predominantly consists of significantly altered genes. In other words, active subnetworks define distinct disease-associated sets of interacting genes, whether discovered through the original analysis or discovered because of being in interaction with a significant gene.
These filtered lists of active subnetworks are then used for enrichment analyses, i.e., using the genes in each of the active subnetworks, the significantly enriched terms (pathways/gene sets) are identified. Enriched terms with adjusted p-values larger than the given threshold are discarded, and the lowest adjusted p-value (among all active subnetworks) for each term is kept. This process of active subnetwork search + enrichment analyses is repeated for a selected number of iterations, performed in parallel. Over all iterations, the lowest and the highest adjusted p-values, and the number of occurrences among all iterations are reported for each significantly enriched term.
This workflow can be run using the function run_pathfindR():
library(pathfindR)
output_df <- run_pathfindR(input_df)This wrapper function performs the active-subnetwork-oriented enrichment analysis, and returns a data frame of enriched terms:
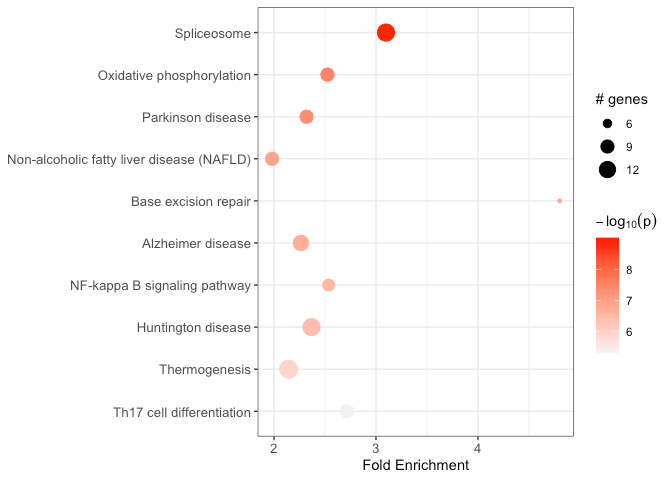
Some useful arguments are:
# set an output directory for saving active subnetworks and creating an HTML report
# (default=NULL, sets a temporary directory)
output_df <- run_pathfindR(input_df, output_dir="/top/secret/results")
# change the gene sets used for analysis (default="KEGG")
output_df <- run_pathfindR(input_df, gene_sets="GO-MF")
# change the PIN for active subnetwork search (default=Biogrid)
output_df <- run_pathfindR(input_df, pin_name_path="IntAct")
# or use an external PIN of your choice
output_df <- run_pathfindR(input_df, pin_name_path="/path/to/my/PIN.sif")
# change the number of iterations (default=10)
output_df <- run_pathfindR(input_df, iterations=25)
# report the non-significant active subnetwork genes (for later analyses)
output_df <- run_pathfindR(input_df, list_active_snw_genes=TRUE)The available PINs are “Biogrid”, “STRING”, “GeneMania”, “IntAct”, “KEGG” and “mmu_STRING”. The available gene sets are “KEGG”, “Reactome”, “BioCarta”, “GO-All”, “GO-BP”, “GO-CC”, “GO-MF”, and “mmu_KEGG”. You also use a custom PIN (see ?return_pin_path) or a custom gene set (see ?fetch_gene_set)
As of the latest development version, pathfindR offers utility functions for obtaining organism-specific PIN data (for now, only BioGRID PINs) and organism-specific gene sets (KEGG and Reactome) data via
get_pin_file()andget_gene_sets_list(), respectively.
Clustering of the Enriched Terms
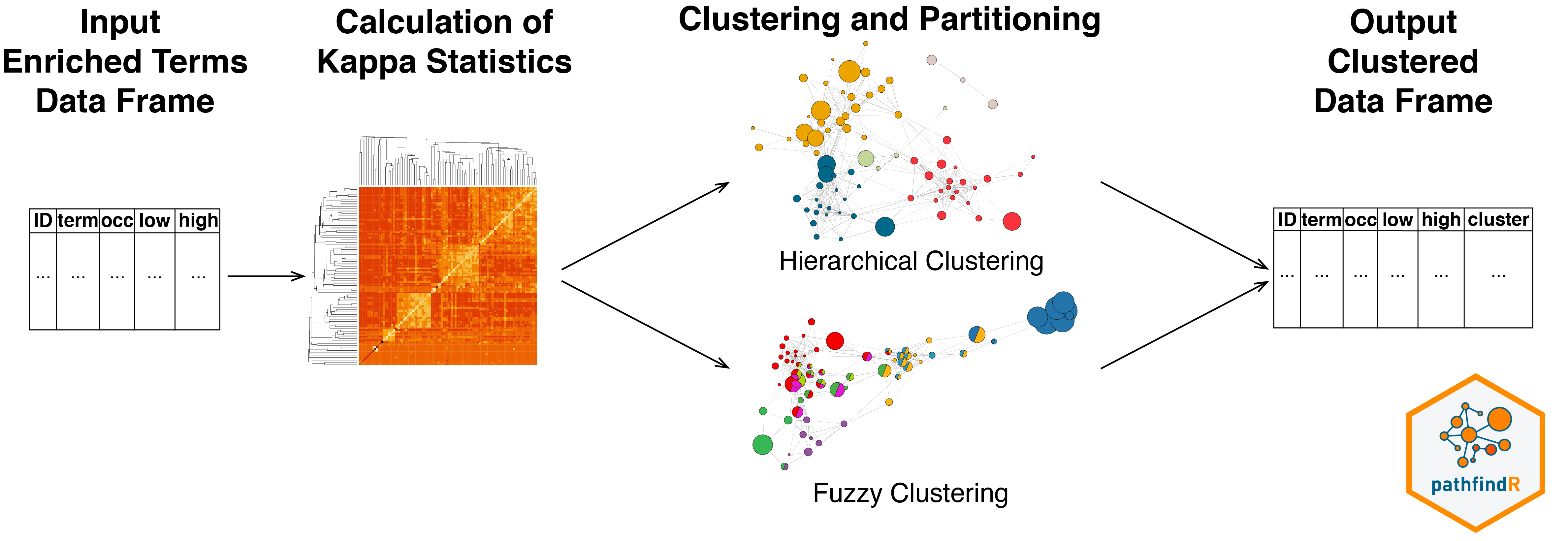 The wrapper function for this workflow is
The wrapper function for this workflow is cluster_enriched_terms().
This workflow first calculates the pairwise kappa statistics between the enriched terms. The function then performs hierarchical clustering (by default), automatically determines the optimal number of clusters by maximizing the average silhouette width and returns a data frame with cluster assignments.
# default settings
clustered_df <- cluster_enriched_terms(output_df)
# display the heatmap of hierarchical clustering
clustered_df <- cluster_enriched_terms(output_df, plot_hmap=TRUE)
# display the dendrogram and automatically-determined clusters
clustered_df <- cluster_enriched_terms(output_df, plot_dend=TRUE)
# change agglomeration method (default="average") for hierarchical clustering
clustered_df <- cluster_enriched_terms(output_df, clu_method="centroid")Alternatively, the fuzzy clustering method (as described in Huang DW, Sherman BT, Tan Q, et al. The DAVID Gene Functional Classification Tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol. 2007;8(9):R183.) can be used:
clustered_df_fuzzy <- cluster_enriched_terms(output_df, method="fuzzy")Visualization of Enrichment Results
Enriched Term Diagrams
For H.sapiens KEGG enrichment analyses, visualize_terms() can be used to generate KEGG pathway diagrams as ggraph (inherits from ggplot) objects (using ggkegg):
input_processed <- input_processing(example_pathfindR_input)
gg_list <- visualize_terms(
result_df = example_pathfindR_output,
input_processed = input_processed,
is_KEGG_result = TRUE
) # this function returns a list of ggraph objects (named by Term ID)
# save one of the plots as PDF image
ggplot2::ggsave(
"hsa04911_diagram.pdf", # path to output, format is determined by extension
gg_list$hsa04911, # what to plot
width = 5 # adjust width
height = 5 # adjust height
) 
Alternatively (i.e., for other types of (non-KEGG) enrichment analyses), an interaction diagram per enriched term can be generated again via visualize_terms(). These diagrams are also returned as ggraph objects:
input_processed <- input_processing(example_pathfindR_input)
gg_list <- visualize_terms(
result_df = example_pathfindR_output,
input_processed = input_processed,
is_KEGG_result = FALSE,
pin_name_path = "Biogrid"
) # this function returns a list of ggraph objects (named by Term ID)
# save one of the plots as PDF image
ggplot2::ggsave(
"diabetic_cardiomyopathy_interactions.pdf", # path to output, format is determined by extension
gg_list$hsa04911, # what to plot
width = 10 # adjust width
height = 6 # adjust height
) 
Term-Gene Heatmap
The function term_gene_heatmap() can visualize the heatmap of enriched terms by the involved input genes. This heatmap allows visual identification of the input genes involved in the enriched terms, and the common or distinct genes between different terms. If the input data frame (same as in run_pathfindR()) is supplied, the tile colors indicate the change values.
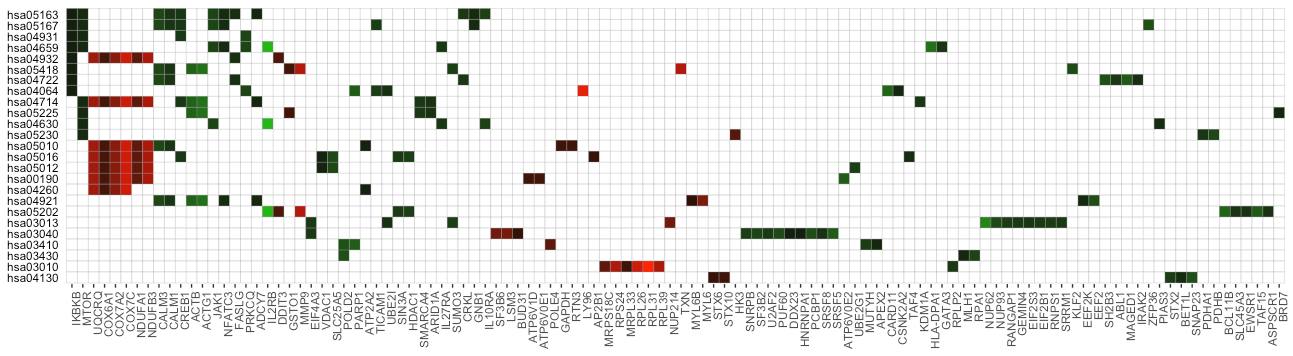
Term-Gene Graph
The function term_gene_graph() (adapted from the Gene-Concept network visualization by the R package enrichplot) can be utilized to visualize which significant genes are involved in the enriched terms. The function creates the term-gene graph, displaying the connections between genes and biological terms (enriched pathways or gene sets). This allows for the investigation of multiple terms to which significant genes are related. The graph also enables the determination of the degree of overlap between the enriched terms by identifying shared and/or distinct significant genes.
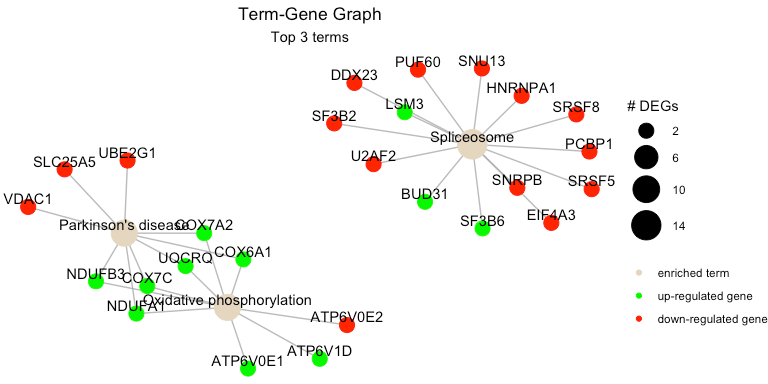
UpSet Plot
UpSet plots are plots of the intersections of sets as a matrix. This function creates a ggplot object of an UpSet plot where the x-axis is the UpSet plot of intersections of enriched terms. By default (i.e., method="heatmap"), the main plot is a heatmap of genes at the corresponding intersections, colored by up-/down-regulation (if genes_df is provided, colored by change values). If method="barplot", the main plot is bar plots of the number of genes at the corresponding intersections. Finally, if method="boxplot" and genes_df is provided, then the main plot displays the boxplots of the genes’ change values at the corresponding intersections.
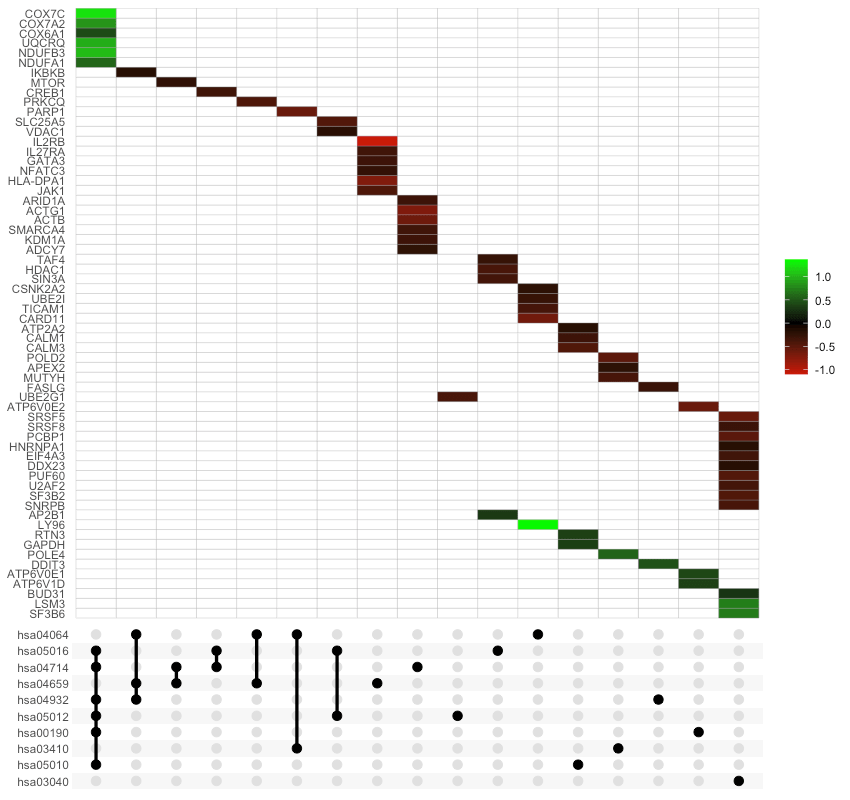
Per Sample Enriched Term Scores
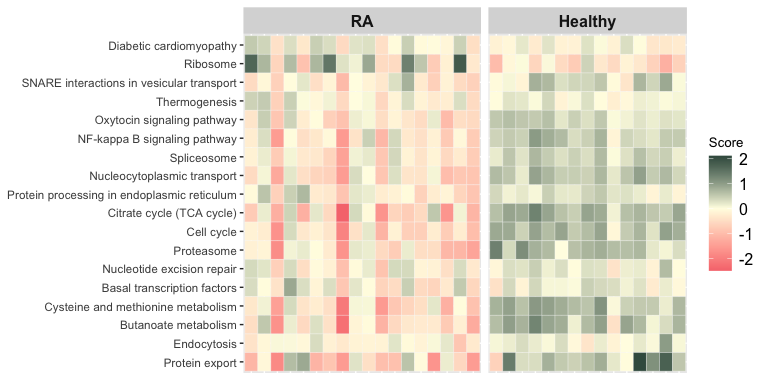
The function score_terms() can be used to calculate the agglomerated z score of each enriched term per sample. This allows the user to examine the scores individually and infer how a term is overall altered (activated or repressed) in a given sample or a group of samples.
Comparison of 2 pathfindR Results
The function combine_pathfindR_results() allows combining two pathfindR analysis results for investigating common and distinct terms between the groups. Below is an example for comparing two different results using rheumatoid arthritis-related data.
combined_df <- combine_pathfindR_results(
result_A=an_output_df,
result_B=another_output_df
)By default, combine_pathfindR_results() plots the term-gene graph for the common terms in the combined results. The function combined_results_graph() can be used to create this graph (using only selected terms etc.) later on.
combined_results_graph(combined_df, selected_terms=c("hsa04144", "hsa04141", "hsa04140"))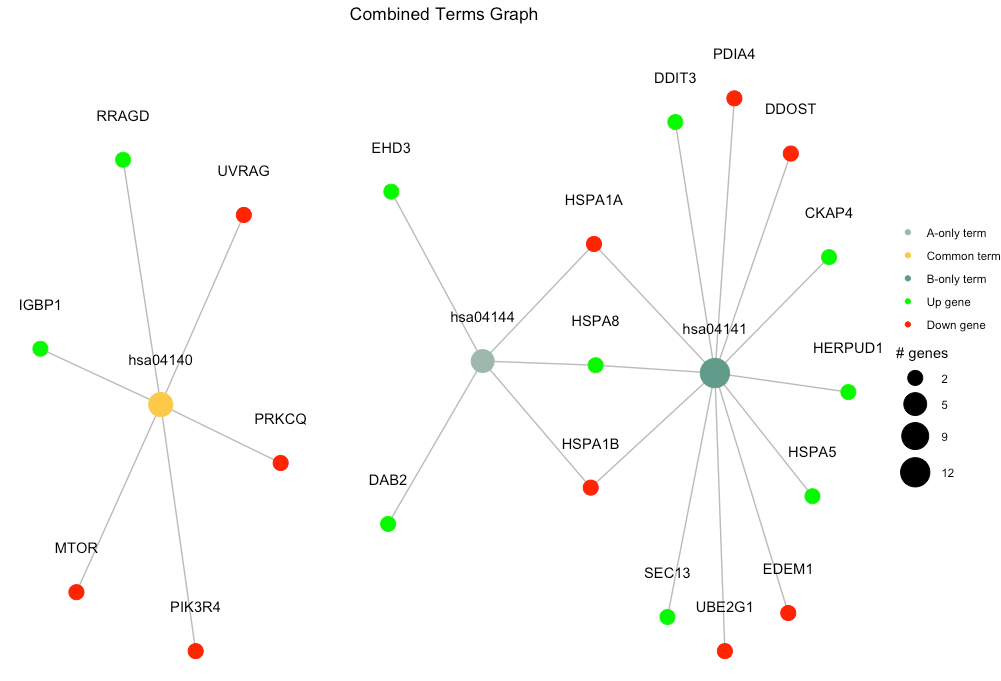
 pathfindR: An R Package for Enrichment Analysis Utilizing Active Subnetworks
pathfindR: An R Package for Enrichment Analysis Utilizing Active Subnetworks
